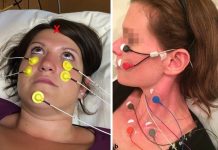
Scientists from the Graduate School of Information Science and Technology at Osaka University used animal location tracking along with artificial intelligence to automatically detect walking behaviors of movement disorders that are shared across species. By automatically removing species-specific features from walking data, the resulting data can be used to better understand neurological disorders that affect movement.
Machine learning algorithms, especially deep learning approaches that use multiple layers of artificial neurons, are well suited for distinguishing between different sources of data. For example, they can determine the species according to the characteristics of its tracks left behind in snow. However, there are times scientists care more about what is the same, rather than what is different, in various datasets. This may be the case when trying to aggregate readings from different types of animals.
Now, a team of scientists led by Osaka University have used machine learning to obtain patterns from locomotion data created by worm, beetle, mouse, and human subjects that were independent of the species. “A central goal of comparative behavioral analysis is to identify human-like behavioral repertoires in animals,” first author Takuya Maekawa explains. This method can help scientists studying human neurological conditions that cause motor dysfunctions, including those resulting from low dopamine levels. Animal motion data would generate much more information; however, the spatial and temporal scales of animal locomotion vary widely among species. This means that the data cannot be directly compared with human behavior. To overcome this, the team designed a neural network with a gradient reversal layer that predicts a) whether or not input locomotion data came from a diseased animal and b) from which species the input data came. From there, the network was trained so that it would fail to predict the species from which the input data was gathered, which resulted in the creation of a network that was incapable of distinguishing between species but capable of identifying specific diseases. This enabled the network to extract locomotion features inherent to the disease.
Their experiments revealed cross-species locomotion features shared by dopamine-deficient worms, mice, and humans. Despite their evolutionary differences, all of these organisms are unable to move while maintaining high speeds. Also, the speed of these animals was found to be unstable when accelerating. Interestingly, these animals exhibit similar movement disorders in the case of dopamine deficiency even though they have different body scales and locomotion methods. While previous studies had shown that dopamine deficiency was associated with movement disorders in all of these species, this research was the first to identify the shared locomotion features caused by this deficiency.
“Our project shows that deep learning can be a powerful tool for extracting knowledge from datasets that appear too different to be compared by human researchers,” author Takahiro Hara says. The team anticipates that this work will be used to find other common features for disorders that impact evolutionarily distant species.